Exploration of SARS-CoV-2 Mpro inhibitors through molecular docking studies
Keywords:
SARS-CoV-2, Mpro inhibitors, Molecular docking, SimulationsAbstract
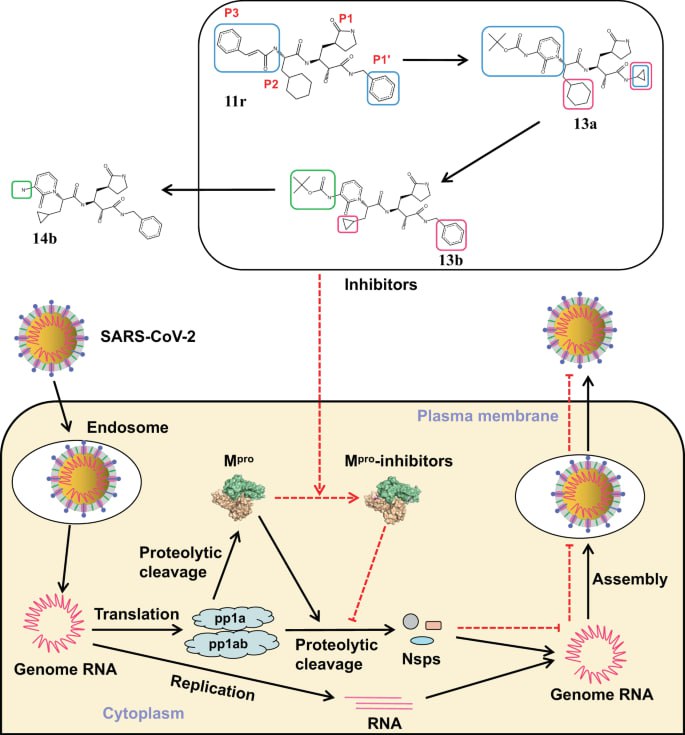
The SARS-CoV-2 main protease (Mpro) is indispensable for viral replication and thus represents a highly attractive therapeutic target. In this study, molecular docking was employed to explore natural bioactive compounds as potential inhibitors of Mpro. The crystal structure of SARS-CoV-2 Mpro (PDB ID: 6LU7) was retrieved from the RCSB Protein Data Bank, and a ligand library of 38 phytochemicals derived from Azadirachta indica, Curcuma longa, Zingiber officinale, Ocimum basilicum, and Panax ginseng was compiled from PubChem. Docking simulations were carried out using AutoDock 4.2 with the Lamarckian Genetic Algorithm to evaluate binding affinities and interaction patterns within the protease’s substrate-binding pockets. Visualization and interaction analyses were performed with LigPlot+ and UCSF Chimera to validate hydrogen bonding, hydrophobic contacts, and potential allosteric effects. Several compounds, particularly flavonoids such as casticin, exhibited strong binding energies and favorable interactions with both the catalytic dyad and allosteric regions of Mpro, suggesting dual mechanisms of inhibition. The integration of phytochemical screening with structure-based docking highlights promising leads for the development of safe, plant-derived antivirals. Collectively, these results provide a computational framework for identifying novel SARS-CoV-2 Mpro inhibitors and contribute to the broader effort of advancing effective therapeutic strategies against COVID-19.
Downloads
Published
Issue
Section
License
Copyright (c) 2025 .

This work is licensed under a Creative Commons Attribution 4.0 International License.